cTWAS_analysis_for_Ulcerative_Colitis
Jing Gu
2023-08-25
Last updated: 2023-08-25
Checks: 6 1
Knit directory: m6A_in_disease_genetics/
This reproducible R Markdown analysis was created with workflowr (version 1.7.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20230331) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Using absolute paths to the files within your workflowr project makes it difficult for you and others to run your code on a different machine. Change the absolute path(s) below to the suggested relative path(s) to make your code more reproducible.
| absolute | relative |
|---|---|
| ~/projects/m6A_in_disease_genetics/code/ctwas/ctwas_config_b37.R | code/ctwas/ctwas_config_b37.R |
| ~/projects/m6A_in_disease_genetics/code/ctwas/qiansheng/locus_plot.R | code/ctwas/qiansheng/locus_plot.R |
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version d0f5634. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .ipynb_checkpoints/
Ignored: analysis/m6A_switch_to_disease_h2g.nb.html
Ignored: data/plots/
Untracked files:
Untracked: HMGCR_locus_gene_tracks.pdf
Untracked: Rplots.pdf
Untracked: analysis/.ipynb_checkpoints/
Untracked: analysis/IBD_E_S_m6A.Rmd
Untracked: analysis/IBD_E_S_m6A_output.Rmd
Untracked: analysis/LDL_E_S_m6A.Rmd
Untracked: analysis/LDL_m6A_output.Rmd
Untracked: analysis/RA_m6A_output.Rmd
Untracked: analysis/WhiteBlood_WholeBlood_E_M.Rmd
Untracked: analysis/identify_m6A_mechanisms_with_finemapping.Rmd
Untracked: analysis/lymph_m6A_output.Rmd
Untracked: analysis/pre_weights_m6AQTL.txt
Untracked: analysis/rbc_E_S_m6A_output.Rmd
Untracked: analysis/rbc_m6A_output.Rmd
Untracked: analysis/summarize_ctwas_m6A_results.Rmd
Untracked: analysis/wbc_E_S_m6A_output.Rmd
Untracked: code/.ipynb_checkpoints/
Untracked: code/all_m6a_sites_with_paired_cisNATs_summary.csv
Untracked: code/annotating_fine-mapped_m6A_QTLs.Rmd
Untracked: code/check_double_strand.ipynb
Untracked: code/check_double_strand_v2.ipynb
Untracked: code/ctwas/
Untracked: code/figure/
Untracked: code/learn_gviz.Rmd
Untracked: code/learn_gviz.html
Untracked: code/learn_gviz.nb.html
Untracked: code/m6AQTL_finemapping.Rmd
Untracked: code/plot_genomic_tracks_gviz.ipynb
Untracked: code/summary_TWAS_coloc_m6A_2023.Rmd
Untracked: code/test_gviz.ipynb
Untracked: code/twas_genes_PP4_0.3_immune_traits_trackplots.pdf
Untracked: data/.ipynb_checkpoints/
Untracked: data/ADCY7_gwas_input.tsv
Untracked: data/ADCY7_qtl_input.tsv
Untracked: data/Allergy_full_coloc.txt
Untracked: data/Asthma_full_coloc.txt
Untracked: data/CAD_full_coloc.txt
Untracked: data/Eosinophil_count_full_coloc.txt
Untracked: data/GSE125377_jointPeakReadCount.txt
Untracked: data/G_list.Rd
Untracked: data/HMGCR_ctwas_dat.Rd
Untracked: data/IBD_full_coloc.txt
Untracked: data/JointPeaks.bed
Untracked: data/Li2022_dsRNAs.xlsx
Untracked: data/Lupus_full_coloc.txt
Untracked: data/RA_full_coloc.txt
Untracked: data/TABLE1_hg19.txt
Untracked: data/TABLE1_hg19.txt.zip
Untracked: data/__MACOSX/
Untracked: data/coloc_blood_traits.csv
Untracked: data/crohns_disease_full_coloc.txt
Untracked: data/ctwas_m6a_joint_top_PIP.txt
Untracked: data/edit_sites_and_GE_neg_correlated.txt
Untracked: data/edit_sites_and_GE_pos_correlated.txt
Untracked: data/features
Untracked: data/human_EERs.csv
Untracked: data/human_EERs.txt
Untracked: data/lymph_full_coloc.txt
Untracked: data/m6A_TWAS_results.csv
Untracked: data/m6a_TWAS_genes.txt
Untracked: data/m6a_joint_calling_peaks.csv
Untracked: data/nasser_2021_ABC_IBD_genes.txt
Untracked: data/nat_sense_pairs.csv
Untracked: data/plt_full_coloc.txt
Untracked: data/rbc_full_coloc.txt
Untracked: data/rdw_full_coloc.txt
Untracked: data/reported_AS_targets_S1.txt
Untracked: data/reported_AS_wanowska.txt
Untracked: data/sig_coloc_results/
Untracked: data/test_locuscomparer.pdf
Untracked: data/ulcerative_colitis_full_coloc.txt
Untracked: data/wbc_full_coloc.txt
Untracked: data/zhao_silver_genes.csv
Untracked: output/.ipynb_checkpoints/
Untracked: output/HMGCR_gene_track_plot.pdf
Untracked: output/HMGCR_locus_plot.pdf
Untracked: output/IBD_DHX38_plot.pdf
Untracked: output/IBD_DHX38_plot_genetrack.pdf
Untracked: output/IBD_IP6K2_plot.pdf
Untracked: output/LDL_PMPCA_plot.pdf
Untracked: output/LDL_TMEM199_plot.pdf
Untracked: output/all_m6a_sites_with_cisNATs.csv
Untracked: output/all_m6a_sites_with_paired_cisNATs_summary.csv
Untracked: output/all_m6a_sites_with_paired_cisNATs_summary_PP40.3.csv
Untracked: output/all_m6a_sites_with_paired_cisNATs_summary_PP40.5.csv
Untracked: output/all_m6a_sites_with_paired_cis_NATs.csv
Untracked: output/asthma_GPR183_plot.pdf
Untracked: output/asthma_GPR183_plot2.pdf
Untracked: output/asthma_GPR183_plot_genetrack.pdf
Untracked: output/asthma_GPR183_plot_genetrack2.pdf
Untracked: output/asthma_SLC25A11_plot_genetrack.pdf
Untracked: output/asthma_ZMAT2_plot.pdf
Untracked: output/asthma_ZMAT2_plot2.pdf
Untracked: output/asthma_ZMAT2_plot_genetrack.pdf
Untracked: output/asthma_ZMAT2_plot_genetrack2.pdf
Untracked: output/ctwas_locus_plot_output_data.Rd
Untracked: output/ctwas_m6a_joint_m6a_PIP_0.8.csv
Untracked: output/ctwas_m6a_joint_m6a_PIP_0.8.txt
Untracked: output/ctwas_m6a_joint_pip_above_0.6.RData
Untracked: output/eosinoph_MEPCE_plot.pdf
Untracked: output/eosinoph_PTK2B_plot.pdf
Untracked: output/eosinoph_PTK2B_plot_genetrack.pdf
Untracked: output/eosinoph_SH2D3C_plot.pdf
Untracked: output/eosinoph_SLC43A3_plot.pdf
Untracked: output/eosinoph_ZER1_plot.pdf
Untracked: output/eosinoph_ZER1_plot_genetrack.pdf
Untracked: output/fine_mapped_m6AQTLs_TWAS_genes_highPP4.rds
Untracked: output/gene_summary.csv
Untracked: output/immune_related_m6A_targets.csv
Untracked: output/lupus_MIR210HG_IRF7_plot.pdf
Untracked: output/lupus_MIR210HG_IRF7_plot_genetrack.pdf
Untracked: output/lupus_MIR210HG_plot.pdf
Untracked: output/lupus_MIR210HG_plot_genetrack.pdf
Untracked: output/lymph_ADCY7_BRD7_plot.pdf
Untracked: output/lymph_ADCY7_BRD7_plot_genetrack.pdf
Untracked: output/lymph_ADCY7_NOD2_plot.pdf
Untracked: output/lymph_ADCY7_NOD2_plot_genetrack.pdf
Untracked: output/lymph_ADCY7_plot.pdf
Untracked: output/lymph_BAZ1B_plot.pdf
Untracked: output/lymph_EPC1_plot.pdf
Untracked: output/lymph_LAMTOR4_plot.pdf
Untracked: output/lymph_MDM2_plot.pdf
Untracked: output/lymph_RANGAP1_plot.pdf
Untracked: output/lymph_TAP2_plot.pdf
Untracked: output/lymph_THEMIS2_plot.pdf
Untracked: output/lymph_ZKSCAN5_plot.pdf
Untracked: output/lymph_ZSCAN25_plot.pdf
Untracked: output/m6aQTL_dsRNAs_PPP2R3C_PRORP.pdf
Untracked: output/m6a_QTL_genes.csv
Untracked: output/m6a_genes_PIP_0.6_blood_immune.csv
Untracked: output/m6a_genes_PIP_0.6_blood_immune.txt
Untracked: output/m6a_peaks_nearby_dsRNAs.csv
Untracked: output/m6a_sites_near_all_dsRNAs_twas.csv
Untracked: output/m6a_sites_near_dsRNAs_coloc.csv
Untracked: output/m6a_sites_near_dsRNAs_twas.csv
Untracked: output/m6a_sites_near_dsRNAs_twas_summary.csv
Untracked: output/m6a_sites_overlapping_NAT_twas.csv
Untracked: output/m6a_sites_overlapping_dsRNAs_coloc.csv
Untracked: output/m6a_sites_overlapping_dsRNAs_twas.csv
Untracked: output/m6a_sites_overlapping_dsRegions.csv
Untracked: output/m6a_sites_overlapping_dsRegions_coloc.csv
Untracked: output/negatively_correlated_genes.txt
Untracked: output/platelet_AKAP8_plot.pdf
Untracked: output/platelet_C17orf62_plot.pdf
Untracked: output/platelet_GSDMD_plot.pdf
Untracked: output/platelet_PHF11_plot.pdf
Untracked: output/platelet_SLC25A11_plot.pdf
Untracked: output/platelet_TAOK1_plot.pdf
Untracked: output/platelet_THEMIS2_plot.pdf
Untracked: output/platelet_THEMIS2_plot_genetrack.pdf
Untracked: output/platelet_TRAF2_plot.pdf
Untracked: output/platelet_UBE2G2_plot.pdf
Untracked: output/platelet_ZSCAN25_plot.pdf
Untracked: output/postively_correlated_genes.txt
Untracked: output/rs1806261_RABEP1-NUP88_focused_locusview.pdf
Untracked: output/rs1806261_RABEP1-NUP88_locusview.pdf
Untracked: output/rs3177647_MAPKAPK5-AS1-MAPKAPK5_locusview.pdf
Untracked: output/rs3204541_DDX55-EIF2B1_locusview.pdf
Untracked: output/rs7184802_ADCY7-BRD7_locusview.pdf
Untracked: output/rs7184802_ADCY7_locuscompare.pdf
Untracked: output/twas_genes_PP4_0.3_immune_traits_trackplots.pdf
Untracked: output/twas_genes_PP4_0.5_blood_traits_trackplots.pdf
Untracked: output/twas_m6a_sites_with_all_cisNATs.RDS
Untracked: output/twas_m6a_sites_with_cisNATs_range.RDS
Untracked: output/twas_m6a_sites_with_the_nearest_cisNAT.RDS
Untracked: output/wbc_RABEP1_NUP88_plot.pdf
Untracked: output/wbc_RABEP1_NUP88_plot_genetrack.pdf
Untracked: output/wbc_SLC9A3R1_plot.pdf
Untracked: twas_genes_PP4_0.3_immune_traits_trackplots.pdf
Unstaged changes:
Deleted: analysis/learn_ctwas.Rmd
Modified: analysis/lymph_m6A_output_hg19.Rmd
Modified: analysis/m6A_switch_to_disease_h2g.Rmd
Modified: analysis/rbc_m6A_output_hg19.Rmd
Modified: analysis/wbc_m6A_output.Rmd
Modified: analysis/wbc_m6A_output_hg19.Rmd
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown (analysis/UC_m6A_output_hg19.Rmd)
and HTML (docs/UC_m6A_output_hg19.html) files. If you’ve
configured a remote Git repository (see ?wflow_git_remote),
click on the hyperlinks in the table below to view the files as they
were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | d0f5634 | Jing Gu | 2023-08-25 | other traits |
Load ctwas results
# top 1 method
res <- impute_expr_z(z_snp, weight = weight, ld_R_dir = ld_R_dir,
method = NULL, outputdir = outputdir, outname = outname.e,
harmonize_z = T, harmonize_wgt = T, scale_by_ld_variance=F,
strand_ambig_action_z = "recover",
recover_strand_ambig_wgt = T
# lasso/elastic-net method
res <- impute_expr_z(z_snp, weight = weight, ld_R_dir = ld_R_dir,
method = NULL, outputdir = outputdir, outname = outname.e,
harmonize_z = T, harmonize_wgt = T, scale_by_ld_variance=F,
strand_ambig_action_z = "none",
recover_strand_ambig_wgt = FGWAS: UK Biobank GWAS summary statistics - European individuals
Weights: FUSION weights using top1, lasso, or elastic-net models were converted into PredictDB format and were not needed to do scaling when running ctwas.
Check convergence of parameters
cTWAS analysis on m6A alone
Joint analysis of expression, splicing and m6A
[1] "Check convergence for the lasso model when jointly analyzing expression, splicing and m6A:"
[1] "Table of group size before/after matching with UKBB SNPs:"
SNP eQTL sQTL m6AQTL
prior_group_size 9.324e+06 2005.0000 2191.0000 918.0000
group_size 3.133e+06 1806.0000 1921.0000 797.0000
percent_of_overlaps 3.360e-01 0.9007 0.8768 0.8682
SNP eQTL sQTL m6AQTL
estimated_group_prior 0.0003549 0.018832 1.618e-03 1.305e-03
estimated_group_prior_var 1.8128614 2.219582 9.458e-01 1.405e+00
estimated_group_pve 0.0043529 0.000163 6.347e-06 3.157e-06
attributable_group_pve 0.9618719 0.036028 1.403e-03 6.976e-04$lasso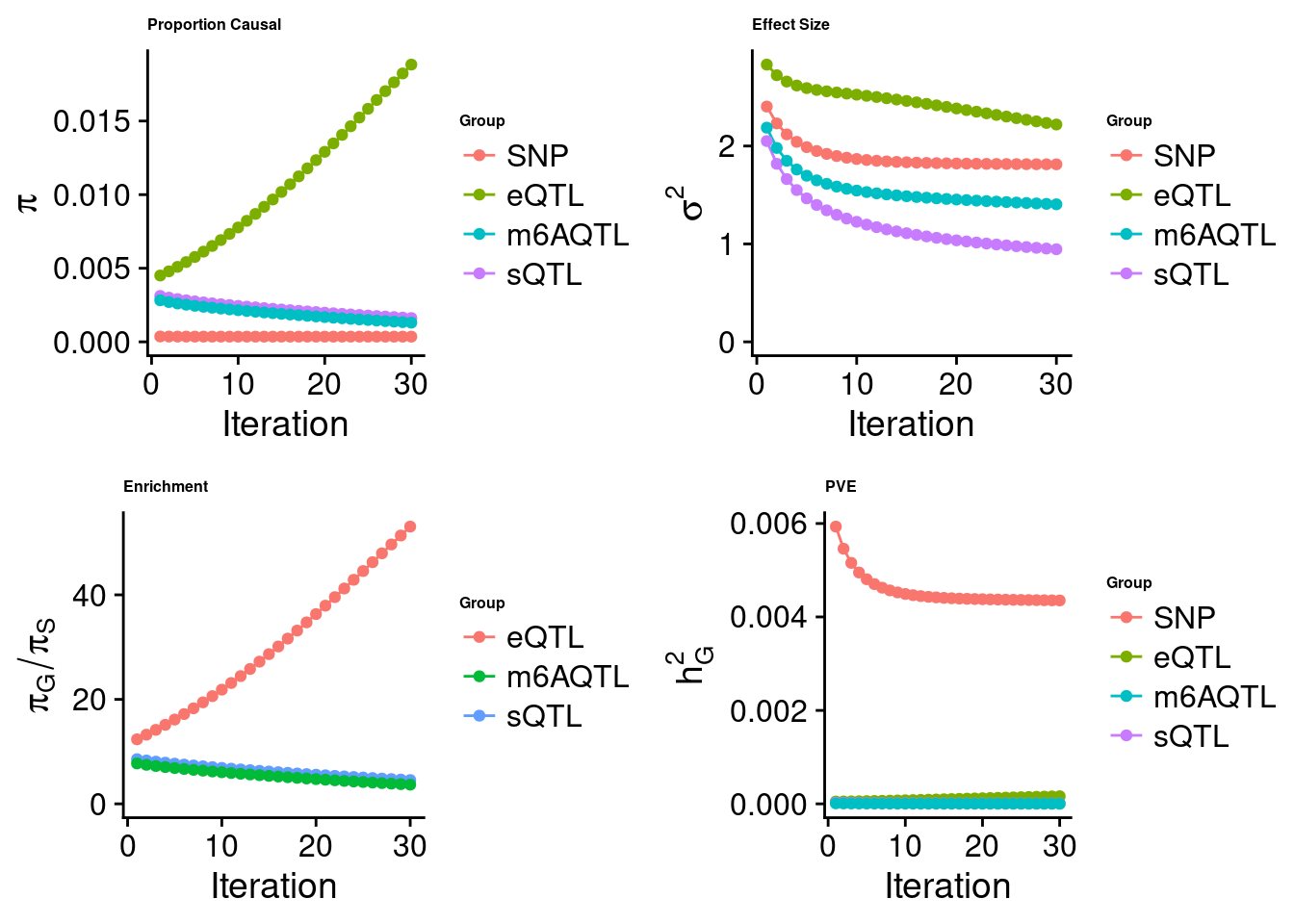
cTWAS results for individual analysis with m6A
top1 model
Summing up PIPs for m6A peaks located in the same gene
Top m6A PIPs by genes
cTWAS results for joint analysis using a lasso model
Top m6A modification pip
Top expression/splicing/m6A units
For m6A or splicing QTLs, they are assigned to the nearest genes (m6A needs to be confirmed with Kevin).
Top SNPs or genes with PIP > 0.6
$eQTL
[1] genename susie_pip group region_tag
<0 rows> (or 0-length row.names)
$m6AQTL
[1] genename susie_pip group region_tag
<0 rows> (or 0-length row.names)
$sQTL
[1] genename susie_pip group region_tag
<0 rows> (or 0-length row.names)Top m6A modification pip
genename region_tag susie_pip z
1 ICOSLG 21_23 0.05737 3.697
2 FUT10 8_32 0.03381 -2.826
3 EIF3B 7_4 0.02885 3.086
4 SORD 15_17 0.02603 2.533
5 KIF21B 1_103 0.02281 -4.045
6 MEF2A 15_48 0.02182 2.437
7 SCIMP 17_5 0.02146 -2.513
8 ZC3H12D 6_97 0.01956 -2.935
9 SLC25A25-AS1 9_66 0.01882 -2.456
10 TRIT1 1_25 0.01792 -2.473Summing up PIPs for m6A peaks located in the same gene
Top 10 m6A PIPs by genes
# A tibble: 723 × 2
genename total_susie_pip
<chr> <dbl>
1 ICOSLG 0.0669
2 FUT10 0.0338
3 EIF3B 0.0289
4 SORD 0.0260
5 KIF21B 0.0228
6 PARP14 0.0223
7 MEF2A 0.0218
8 SCIMP 0.0215
9 SUGP2 0.0207
10 ZC3H12D 0.0196
# ℹ 713 more rowsTop splicing PIPs
peak_id genename pos region_tag susie_pip z
1 chr14:75991531-76024457 BATF 75937022 14_34 0.03772 -3.003
2 chr7:25163746-25164309 CYCS 25115972 7_22 0.03171 2.843
3 chr2:10583439-10583616 ODC1 10666469 2_7 0.02680 -2.576
4 chr20:33163050-33176257 PIGU 33195608 20_21 0.02548 -3.614
5 chr8:99126053-99129260 RIDA 99125841 8_67 0.02481 2.515
6 chr14:90865431-90866409 CALM1 90817732 14_45 0.02404 2.511
7 chr19:50925802-50926862 SPIB 50926742 19_34 0.02387 -3.444
8 chr20:43514527-43530172 YWHAB 43432189 20_28 0.02307 -3.105
9 chr20:43516383-43530172 YWHAB 43514337 20_28 0.02300 3.102
10 chr17:5124645-5138030 SCIMP 5124462 17_5 0.02296 2.559Summing up PIPs for spliced introns located in the same gene
Top 10 splicing PIPs by genes
# A tibble: 10 × 2
genename total_susie_pip
<chr> <dbl>
1 IMMP1L 0.116
2 MCOLN2 0.0914
3 SYNCRIP 0.0906
4 SP140 0.0904
5 WARS1 0.0869
6 ALDH3A2 0.0804
7 IFI44L 0.0717
8 SLAMF7 0.0713
9 LGALS9 0.0672
10 LYRM1 0.0645Top genes by combined PIP
genename combined_pip expression_pip splicing_pip m6A_pip
871 ENSG00000227039 0.379 0.3785 0.000000 0.000000
1828 NELFCD 0.372 0.3671 0.000000 0.004567
707 DSE 0.364 0.3640 0.000000 0.000000
2623 SYNGR1 0.360 0.3599 0.000000 0.000000
265 BCAS4 0.338 0.3334 0.005077 0.000000
2987 YWHAB 0.325 0.2715 0.053020 0.000000
58 ACAD8 0.312 0.3116 0.000000 0.000000
1716 MRPL42 0.299 0.2993 0.000000 0.000000
1970 PCNX4 0.285 0.2852 0.000000 0.000000
2572 SSR3 0.282 0.2370 0.044779 0.000000
3015 ZKSCAN3 0.274 0.2744 0.000000 0.000000
1413 IL10RB 0.271 0.2616 0.000000 0.009632
2978 YBEY 0.270 0.2617 0.008367 0.000000
1538 LIN9 0.260 0.2603 0.000000 0.000000
1921 ORC5 0.260 0.2540 0.006287 0.000000
2019 PIGX 0.255 0.2554 0.000000 0.000000
911 ENSG00000235560 0.252 0.2525 0.000000 0.000000
938 ENSG00000243150 0.238 0.2382 0.000000 0.000000
1109 ENSG00000272947 0.233 0.2329 0.000000 0.000000
2411 SFXN2 0.233 0.2328 0.000000 0.000000
region_tag
871 21_23
1828 20_34
707 6_77
2623 22_16
265 20_31
2987 20_28
58 11_83
1716 12_55
1970 14_27
2572 3_97
3015 6_22
1413 21_14
2978 21_24
1538 1_117
1921 7_64
2019 3_121
911 16_24
938 3_98
1109 3_77
2411 10_66Loading required package: gridWarning: replacing previous import 'utils::download.file' by
'restfulr::download.file' when loading 'rtracklayer'Locus plots for specific examples
R version 4.2.0 (2022-04-22)
Platform: x86_64-pc-linux-gnu (64-bit)
Running under: CentOS Linux 7 (Core)
Matrix products: default
BLAS/LAPACK: /software/openblas-0.3.13-el7-x86_64/lib/libopenblas_haswellp-r0.3.13.so
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C LC_TIME=C
[4] LC_COLLATE=C LC_MONETARY=C LC_MESSAGES=C
[7] LC_PAPER=C LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C LC_IDENTIFICATION=C
attached base packages:
[1] grid stats4 stats graphics grDevices utils datasets
[8] methods base
other attached packages:
[1] biomaRt_2.52.0 Gviz_1.40.1 cowplot_1.1.1
[4] ggplot2_3.4.3 GenomicRanges_1.48.0 GenomeInfoDb_1.32.2
[7] IRanges_2.30.1 S4Vectors_0.34.0 BiocGenerics_0.42.0
[10] ctwas_0.1.38 dplyr_1.1.2 workflowr_1.7.0
loaded via a namespace (and not attached):
[1] colorspace_2.1-0 deldir_1.0-6
[3] rjson_0.2.21 rprojroot_2.0.3
[5] biovizBase_1.44.0 htmlTable_2.4.0
[7] XVector_0.36.0 base64enc_0.1-3
[9] fs_1.6.3 dichromat_2.0-0.1
[11] rstudioapi_0.15.0 farver_2.1.1
[13] bit64_4.0.5 AnnotationDbi_1.58.0
[15] fansi_1.0.4 xml2_1.3.3
[17] codetools_0.2-18 logging_0.10-108
[19] cachem_1.0.8 knitr_1.39
[21] Formula_1.2-4 jsonlite_1.8.7
[23] Rsamtools_2.12.0 cluster_2.1.3
[25] dbplyr_2.3.3 png_0.1-7
[27] compiler_4.2.0 httr_1.4.7
[29] backports_1.4.1 lazyeval_0.2.2
[31] Matrix_1.6-1 fastmap_1.1.1
[33] cli_3.6.1 later_1.3.0
[35] htmltools_0.5.2 prettyunits_1.1.1
[37] tools_4.2.0 gtable_0.3.3
[39] glue_1.6.2 GenomeInfoDbData_1.2.8
[41] rappdirs_0.3.3 Rcpp_1.0.11
[43] Biobase_2.56.0 jquerylib_0.1.4
[45] vctrs_0.6.3 Biostrings_2.64.0
[47] rtracklayer_1.56.0 iterators_1.0.14
[49] xfun_0.30 stringr_1.5.0
[51] ps_1.7.0 lifecycle_1.0.3
[53] ensembldb_2.20.2 restfulr_0.0.14
[55] XML_3.99-0.14 getPass_0.2-2
[57] zlibbioc_1.42.0 scales_1.2.1
[59] BSgenome_1.64.0 VariantAnnotation_1.42.1
[61] ProtGenerics_1.28.0 hms_1.1.3
[63] promises_1.2.0.1 MatrixGenerics_1.8.0
[65] parallel_4.2.0 SummarizedExperiment_1.26.1
[67] AnnotationFilter_1.20.0 RColorBrewer_1.1-3
[69] yaml_2.3.5 curl_5.0.2
[71] memoise_2.0.1 gridExtra_2.3
[73] sass_0.4.1 rpart_4.1.16
[75] latticeExtra_0.6-30 stringi_1.7.12
[77] RSQLite_2.3.1 highr_0.9
[79] BiocIO_1.6.0 foreach_1.5.2
[81] checkmate_2.1.0 GenomicFeatures_1.48.4
[83] filelock_1.0.2 BiocParallel_1.30.3
[85] rlang_1.1.1 pkgconfig_2.0.3
[87] matrixStats_0.62.0 bitops_1.0-7
[89] evaluate_0.15 lattice_0.20-45
[91] htmlwidgets_1.5.4 GenomicAlignments_1.32.0
[93] labeling_0.4.2 bit_4.0.5
[95] processx_3.8.0 tidyselect_1.2.0
[97] magrittr_2.0.3 R6_2.5.1
[99] generics_0.1.3 Hmisc_5.1-0
[101] DelayedArray_0.22.0 DBI_1.1.3
[103] pgenlibr_0.3.6 pillar_1.9.0
[105] whisker_0.4 foreign_0.8-82
[107] withr_2.5.0 KEGGREST_1.36.2
[109] RCurl_1.98-1.7 nnet_7.3-17
[111] tibble_3.2.1 crayon_1.5.2
[113] interp_1.1-4 utf8_1.2.3
[115] BiocFileCache_2.4.0 rmarkdown_2.14
[117] jpeg_0.1-10 progress_1.2.2
[119] data.table_1.14.8 blob_1.2.4
[121] callr_3.7.3 git2r_0.30.1
[123] digest_0.6.33 httpuv_1.6.5
[125] munsell_0.5.0 bslib_0.3.1